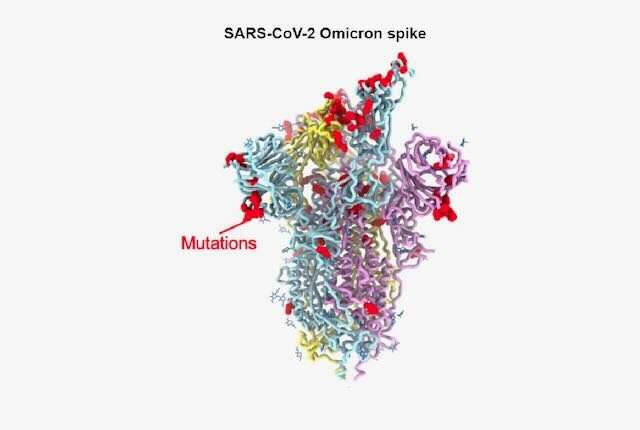
An international team of scientists has determined the precise structural changes in the spike protein of the omicron variant. Their observations explain how the virus is able to evade antibodies against previous variants and still remain highly infectious.
“The findings provide a blueprint that researchers can use to design new countermeasures, whether they be vaccines or therapeutics, against omicron and other coronavirus variants that may emerge,” said David Veesler, investigator with the Howard Hughes Medical Institute and associate professor of biochemistry at the University of Washington School of Medicine in Seattle. He led the research effort with Gyorgy Snell from Vir Biotechnology, Inc. in San Francisco.
The researchers report their findings in the journal Science.
Matthew McCallum, a postdoctoral fellow in Veesler’s lab, and Nadine Czudnochowski, a Vir Biotechnology scientist, were lead authors on the paper.
The omicron variant, which was first identified in November 2021 in South Africa, is causing a surge of infections around the world. In addition to being highly infectious, the variant can evade antibodies against earlier variants leading to breakthrough infections among those who have been vaccinated and those who have been infected previously.
The infectiousness of the virus is thought to be at least in part due to the large number of mutations in the amino acid sequences of the virus’s spike protein. The virus uses the spike protein to latch on to and enter the cells it infects. The omicron spike protein has 37 mutations that distinguish it from the first SARS-CoV-2 isolates in 2020.
Previous research by Veesler and colleagues have shown that antibodies generated by the six most commonly used vaccines, and all but one of monoclonal antibodies currently used to treat infections, have a reduced or abrogated ability to neutralize omicron.
But many of the mutations in the variant affect the structure of the region of the spike protein that is responsible for attaching to and entering cells, a region called the receptor binding domain, and many expected the resulting changes in the receptor binding domain structure might impair the ability of the variant to bind to its target on cells. This target is protein called angiotensin converting enzyme-2, or ACE2. However, in their study, Veesler and his colleagues found that the changes had actually increased the ability of the receptor binding domain to bind to ACE2 by 2.4-fold.
To understand how omicron accumulated so many mutations while retaining efficient interactions with the host receptor ACE2, Veesler and his colleagues used cryo-electron microscopic and X-ray crystallographic studies to unveil the 3-D organization of the omicron spike protein. The approach allowed them to achieve a resolution of about 3 angstroms. At this resolution, it was possible to discern the shape of individual amino acid building blocks that make up the spike protein. The researchers also determined how the structural changes in the spike protein affected the ability of antibodies effective against previous variants to bind to Omicron.
Using these techniques, the scientists reveal how the mutations changed how the protein interacts with antibodies so that the ability of almost all monoclonal antibodies against it is reduced, while, at the same time the ability of the spike receptor-binding domain to bind ACE2 is enhanced. The overall effect has been to make it possible for the receptor binding domain to evade antibodies targeting it and to bind to ACE2 even more tightly.
The findings demonstrate what a formidable opponent SARS-CoV-2 is, says Veesler.
“This virus has incredible plasticity: It can change a lot and still maintain all the functions it needs to infect and replicate,” he noted. “And it’s almost guaranteed omicron is not the last variant we’re going to see.”
The goal going forward should be to focus on and identify additional regions on the spike protein that cannot be changed without causing the protein to lose function, Veesler said. Because of their importance, these areas tend to remain conserved even as other parts of the protein mutates.
Source: Read Full Article